Classification of Skin Lesions with Deep Features Based on SqueezeNet Using Machine Learning Methods
DOI:
https://doi.org/10.58190/imiens.2026.168Keywords:
Skin Lesion, SqueezeNet, Skin Image Analysis, ISIC 2019 Dataset, Image Processing, ClassificationAbstract
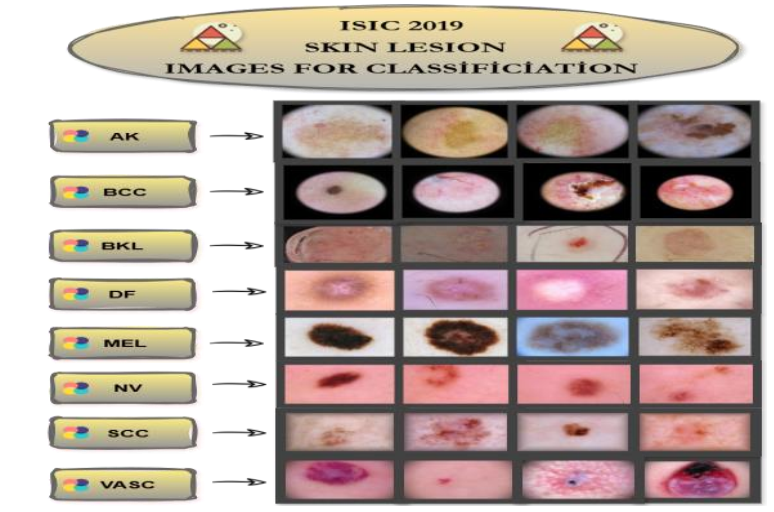
Early and accurate identification of skin lesions is one of the most important factors that improves treatment success in dermatological diseases. Deep learning and image processing algorithms can provide promising results in diagnostic processes by learning complex texture, pigment, and structural features that are difficult for the human eye to distinguish. In this study, a hybrid classification approach based on deep feature extraction and machine learning is proposed. A total of 1,763 dermoscopic images, including eight lesion classes, selected from the ISIC 2019 skin lesion dataset were used. Deep features were extracted using a pretrained SqueezeNet model and analyzed using a 10-fold cross-validation method. The extracted features were evaluated using Artificial Neural Network (ANN), K-Nearest Neighbor (KNN), Support Vector Machine (SVM), and Logistic Regression (LR) classifiers. Experimental results show that the proposed model achieved accuracy values ranging from 96.7% to 89.6% and F1-score values ranging from 96.6% to 89.5%, successfully classifying skin lesions. The KNN model showed relatively lower discrimination capability compared to the other models. All models achieved 100% accuracy in the BCC (Basal Cell Carcinoma) class, while ANN and LR models also classified the SCC (Squamous Cell Carcinoma) class without error. Among all classes, the lowest performance was observed in the DF (Dermatofibroma) class. This study demonstrates that high classification performance can be achieved in multi-class (8 classes) dermatological analysis using lightweight architectures without requiring computationally heavy deep learning models. However, further validation on larger and more diverse datasets is required before real-world clinical applicability can be considered.
Downloads
References
[1] H. Sung et al., "Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries," CA: A Cancer Journal for Clinicians, vol. 71, no. 3, pp. 209–249, 2021, doi: 10.3322/caac.21660.
[2] Iandola, F.N., et al., SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and< 0.5 MB model size. arXiv preprint arXiv:1602.07360, 2016.
[3] W. Gouda et al., "Detection of skin cancer based on skin lesion images using deep learning," Healthcare, vol. 10, no. 6, p. 1057, 2022, doi: 10.3390/healthcare10061057.
[4] N. Nigar et al., "A deep learning approach based on explainable artificial intelligence for skin lesion classification," IEEE Access, vol. 10, pp. 113715–113725, 2022, doi: 10.1109/ACCESS.2022.3212277.
[5] I. Iqbal et al., "Automated multi-class classification of skin lesions through deep convolutional neural network with dermoscopic images," Computerized Medical Imaging and Graphics, vol. 88, p. 101843, 2021, doi: 10.1016/j.compmedimag.2020.101843.
[6] M. A. Kassem et al., "Machine learning and deep learning methods for skin lesion classification and diagnosis: A systematic review," Diagnostics, vol. 11, no. 8, p. 1390, 2021, doi: 10.3390/diagnostics11081390.
[7] M. Goyal et al., "Artificial intelligence-based image classification methods for diagnosis of skin cancer: Challenges and opportunities," Computers in Biology and Medicine, vol. 127, p. 104065, 2020, doi: 10.1016/j.compbiomed.2020.104065.
[8] B. Shetty et al., "Skin lesion classification of dermoscopic images using machine learning and convolutional neural network," Scientific Reports, vol. 12, no. 1, p. 18134, 2022, doi: 10.1038/s41598-022-22644-9.
[9] F. Olayah et al., "AI techniques of dermoscopy image analysis for the early detection of skin lesions based on combined CNN features," Diagnostics, vol. 13, no. 7, p. 1314, 2023, doi: 10.3390/diagnostics13071314.
[10] M. Ahammed, M. Al Mamun, and M. S. Uddin, "A machine learning approach for skin disease detection and classification using image segmentation," Healthcare Analytics, vol. 2, p. 100122, 2022, doi: 10.1016/j.health.2022.100122.
[11] Y. Wang et al., "SSD-KD: A self-supervised diverse knowledge distillation method for lightweight skin lesion classification using dermoscopic images," Medical Image Analysis, vol. 84, p. 102693, 2023, doi: 10.1016/j.media.2023.102693.
[12] N. Hameed et al., "Mobile-based skin lesions classification using convolution neural network," 2023.
[13] P. Tschandl, C. Rosendahl, and H. Kittler, "The HAM10000 dataset: A large collection of multi-source dermatoscopic images of common pigmented skin lesions," Scientific Data, vol. 5, p. 180161, 2018, doi: 10.1038/sdata.2018.161.
[14] N. C. Codella et al., "Skin lesion analysis toward melanoma detection: A challenge at the 2017 International Symposium on Biomedical Imaging," in Proc. IEEE ISBI, 2018, pp. 168–172, doi: 10.1109/ISBI.2018.8363547.
[15] M. Combalia et al., "BCN20000: Dermoscopic lesions in the wild," arXiv preprint arXiv:1908.02288, 2019.
[16] ISIC, "ISIC 2019 Skin Lesion Images for Classification," [Online]. Available: https://kaggle.com/datasets/shajinrp/skin-lesion-image.
[17] Y. LeCun et al., "Gradient-based learning applied to document recognition," Proceedings of the IEEE, vol. 86, no. 11, pp. 2278–2324, 1998, doi: 10.1109/5.726791.
[18] T. Cover and P. Hart, "Nearest neighbor pattern classification," IEEE Transactions on Information Theory, vol. 13, no. 1, pp. 21–27, 1967, doi: 10.1109/TIT.1967.1053964.
[19] V. Vapnik, "Support-vector networks," Machine Learning, vol. 20, pp. 273–297, 1995, doi: 10.1007/BF00994018.
[20] D. W. Hosmer, S. Lemeshow, and R. X. Sturdivant, Applied Logistic Regression, 3rd ed. New York: Wiley, 2013, doi: 10.1002/9781118548387.
[21] D. M. Powers, "Evaluation: From precision, recall and F-measure to ROC, informedness, markedness and correlation," arXiv preprint arXiv:2010.16061, 2020.
[22] J. Allgaier and R. Pryss, "Cross-validation visualized: A narrative guide to advanced methods," Machine Learning and Knowledge Extraction, vol. 6, no. 2, pp. 1378–1388, 2024, doi: 10.3390/make6020071.
Downloads
Published
Issue
Section
License
Copyright (c) 2026 Intelligent Methods In Engineering Sciences

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.






